What is a pathogen?
A pathogen is anything that can produce disease. This can be things like be a virus, fungi or bacteria. While most bacteria in food can be harmless or helpful, some can cause problems, like infections. Some bacteria, in small amounts, are harmless to most healthy adults, while others can multiply and spread and people can become ill. Bacteria that cause illness are known as bacterial pathogens.
Foods that are contaminated with bacteria may not look, taste or smell different from foods that are safe to eat. To prevent or limit illness, scientists at the Canadian Food Inspection Agency (CFIA) work to quickly identify these bacterial pathogens in food. The work of CFIA scientists is essential in tracking down the sources of bacterial contamination in food when it happens.
Genomics and DNA barcoding
Genomics helps us understand, interpret, and use DNA to create solutions to problems that can occur in food. The CFIA carries out genomics research to develop technologies that help scientists identify and understand specific pathogens. These technologies provide new ways to diagnose problems, and lead to faster, cheaper solutions.
Using genomics is a huge improvement from the older biochemical techniques that are used to fight food-borne illnesses. This is like the difference between a detective only knowing a suspect's height and rough physical description compared to having the suspect's fingerprints and behaviour profile. This detailed information makes it much easier to identify the source of contamination leading to a food-borne illness.
CFIA scientists can identify the complete DNA sequence of an organism's genome at a single time using a process known as Whole Genome Sequencing (WGS). This can be done in as little as 24 hours. Once an organism's entire genome is known, a short piece of it can be used as a way of identifying that species whenever its DNA is found. That short piece of the genome acts like a barcode – whenever scientists see that piece of DNA, they know without a doubt what species they're dealing with.
Did you know?
Bacteria like E. coli have around 4 million nucleotides. These are the basic structures that make up DNA. When scientists look for a line of DNA to identify an organism, they take a section of around only 700 nucleotides to act as a barcode. This barcode can be used to differentiate one species of bacteria from every other species.
That's only 0.0175 % of its overall DNA!
Pinpointing Food Pathogens
CFIA scientists can use genomic technologies to sequence the genome of a bacteria taken from a food manufacturing site or from food that has made somebody sick, allowing them to accurately detect and analyze food-borne illnesses and how potentially harmful they are to humans.
Biochemical techniques used by scientists can provide basic information about a sample from contaminated food, like whether the pathogen Listeria is present. Comparatively, a bacteria's DNA barcode can reveal information like its exact strain or how resistant it is to antibiotics. Knowing the bacteria's genome can help inspectors trace a specific sample from the food that it contaminated back to specific areas in a factory to identify where problems are coming from.
How whole genome sequencing works
Whole genome sequencing (WGS) is a laboratory procedure that determines the entire genetic structure of an organism in one process, similar to a blueprint for a building. WGS provides a very precise DNA fingerprint that can help link cases of illness or contamination to one another, allowing an outbreak to be detected and solved sooner.
Click on image for larger view
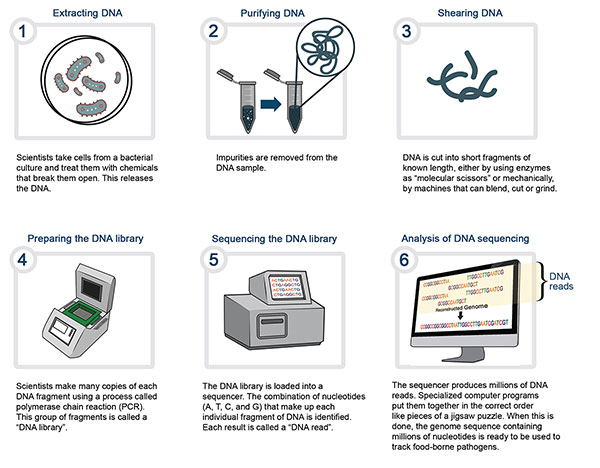
Description for Whole genome sequencing
- Extracting DNA – Scientists take cells from a bacterial culture and treat them with chemicals that break them open. This releases the DNA.
- Purifying DNA – Impurities are removed from the DNA sample.
- Shearing DNA – is cut into short fragments of known length, either by using enzymes as "molecular scissors" or mechanical disruption.
- Preparing the DNA library – Scientists make many copies of each DNA fragment using a process called polymerase chain reaction (PCR). This group of fragments is called a "DNA library."
- Sequencing the DNA library – The DNA library is loaded into a sequencer. The combination of nucleotides (A, T, C, and G) that make up each individual fragment of DNA is identified. Each result is called a "DNA Read."
- Analysis of DNA sequencing – The sequencer produces millions of DNA reads. Specialized computer programs are used to put them together in the correct order like pieces of a jigsaw puzzle. When this is done, the genome sequence containing millions of nucleotides is ready to be used to track food-borne pathogens.
Did you know?
The CFIA has sequenced the entire DNA structure of over 4,000 bacteria that are related to food-borne illnesses, like E. coli, Salmonella, and Listeria.
Following up on problems
When CFIA inspectors identify where problems with bacteria are coming from – for example a certain area of a food processing plant – corrective actions can be identified. If inspectors find the exact same type of bacteria when they come back for a follow-up inspection, the use of this DNA strain-based approach can let them know whether the original problem was not properly addressed, or whether they have an entirely new problem on their hands.
If the new sample of bacteria matches the DNA barcode of the sample they found the first time, this tells the inspectors that the plant may not have fully corrected the problem.
Labs across Canada
The CFIA has DNA sequencing machines in labs across the country. This means sequencing bacteria in food-borne illness outbreaks can be done rapidly – in some cases twice as quickly as older biochemical methods. Analyzing the large amounts of data that come from the CFIA's food microbiology lab's sequencing activities – known as bioinformatics – is done centrally at the CFIA's Ottawa Carling Laboratory.
Bioinformatics software, developed by the CFIA and named GeneSeekr, can identify certain important features in DNA sequencing data. For example, the species or strain of the bacteria, how harmful it is, and markers that indicate whether that bacteria is resistant to antibiotics. The software allows non-scientists to understand complex information quickly and easily to help make decisions in food safety investigations.
Click on image for larger view 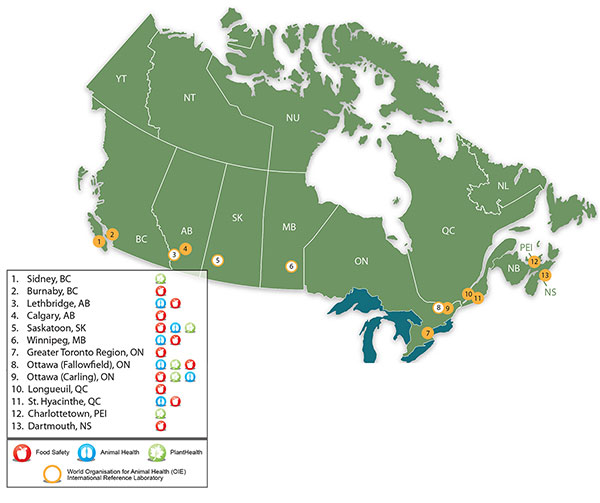
Description of Map showing the location of CFIA laboratories
- Sidney, British Colombia – Plant health
- Burnaby, British Colombia – Food safety
- Lethbridge, Alberta – Food safety, animal health, World Organisation for Animal Health international reference laboratory
- Calgary, Alberta – Food safety
- Saskatoon, Saskatoon – Food safety, animal health, plant health, World Organisation for Animal Health international reference laboratory
- Winnipeg, Manitoba – Food safety, animal health, plant health, World Organisation for Animal Health international reference laboratory
- Greater Toronto Region, Ontario – Food safety
- Ottawa (Fallowfield), Ontario – Food safety, animal health, plant health, World Organisation for Animal Health international reference laboratory
- Ottawa (Carling), Ontario – Food safety, animal health, plant health
- Longueil, Quebec – Food safety
- St-Hyacinthe, Quebec – Food safety, animal health, plant health
- Charlottetown, Prince Edward Island – Plant health
- Dartmouth, Nova Scotia – Food safety
